RADICAL-Pilot
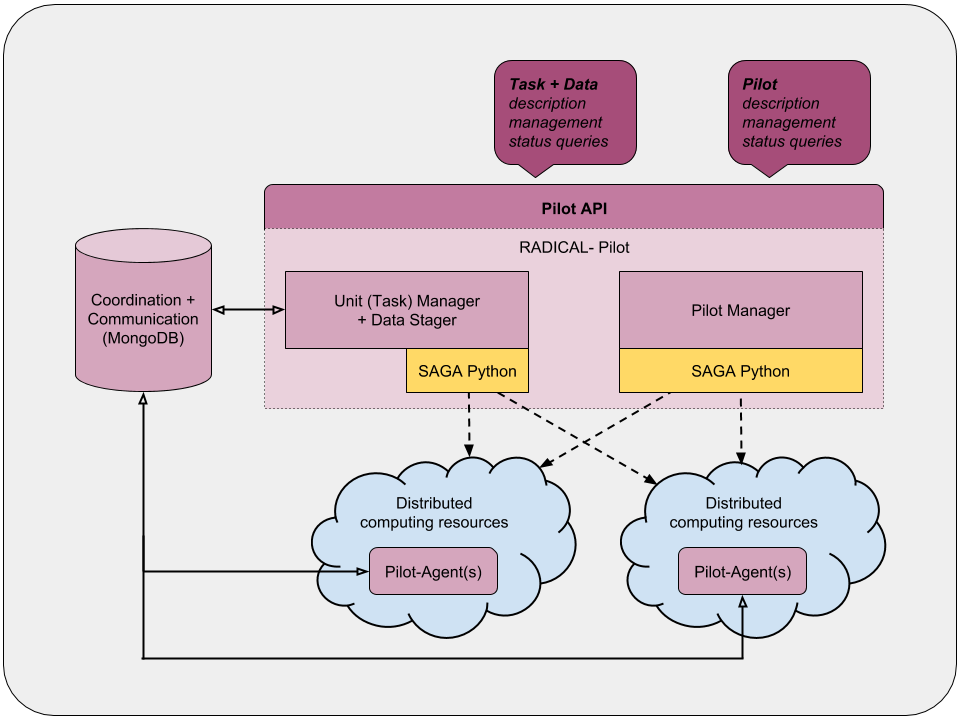
RADICAL-Pilot uses pilots in order to achieve the scalable execution of large numbers of tasks. A pilot is a job submitted to a machine in order to acquire exclusive use of a chunk of its resources. Once the pilot becomes active, it can execute the tasks specified by the user. Instead of having each task waiting on a machine to be executed, the user only has to wait once with a pilot! RADICAL-Pilot allows users to specify resources in the form of pilots and their tasks separately so the user can wait once and run everything.
Basic Example Getting Started More Info Documentation Github Project